

We present DiffInfinite, a hierarchical diffusion model that generates arbitrarily large histological images while preserving long-range correlation structural information. Our approach first generates synthetic segmentation masks, subsequently used as conditions for the high-fidelity generative diffusion process. The proposed sampling method can be scaled up to any desired image size while only requiring small patches for fast training. Moreover, it can be parallelized more efficiently than previous large-content generation methods while avoiding tiling artefacts. The training leverages classifier-free guidance to augment a small, sparsely annotated dataset with unlabelled data. Our method alleviates unique challenges in histopathological imaging practice: large-scale information, costly manual annotation, and protective data handling. The biological plausibility of DiffInfinite data is validated in a survey by ten experienced pathologists as well as a downstream segmentation task. Furthermore, the model scores strongly on anti-copying metrics which is beneficial for the protection of patient data.
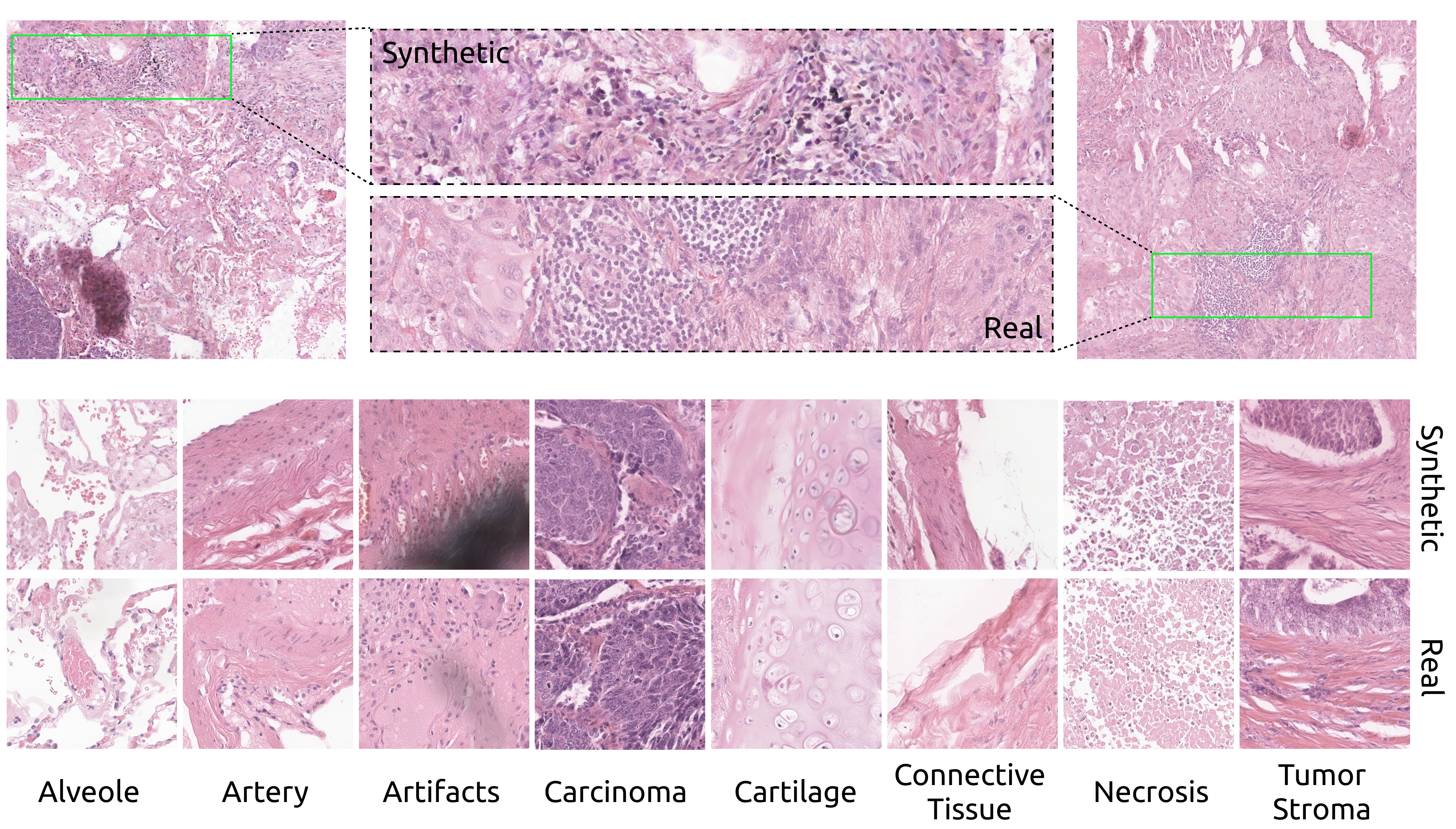
Comparison between real and synthetic images (2048x2048 pixels)
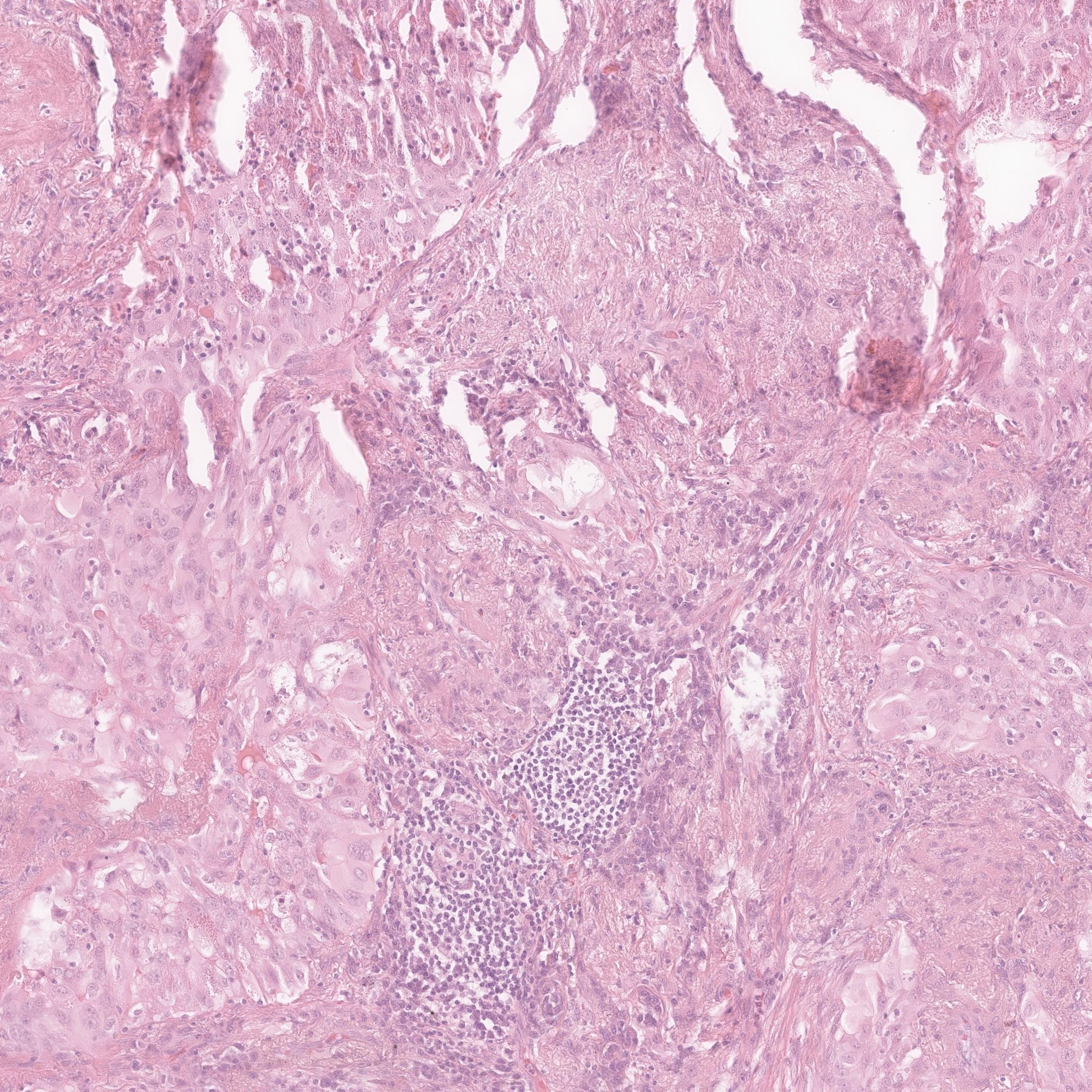
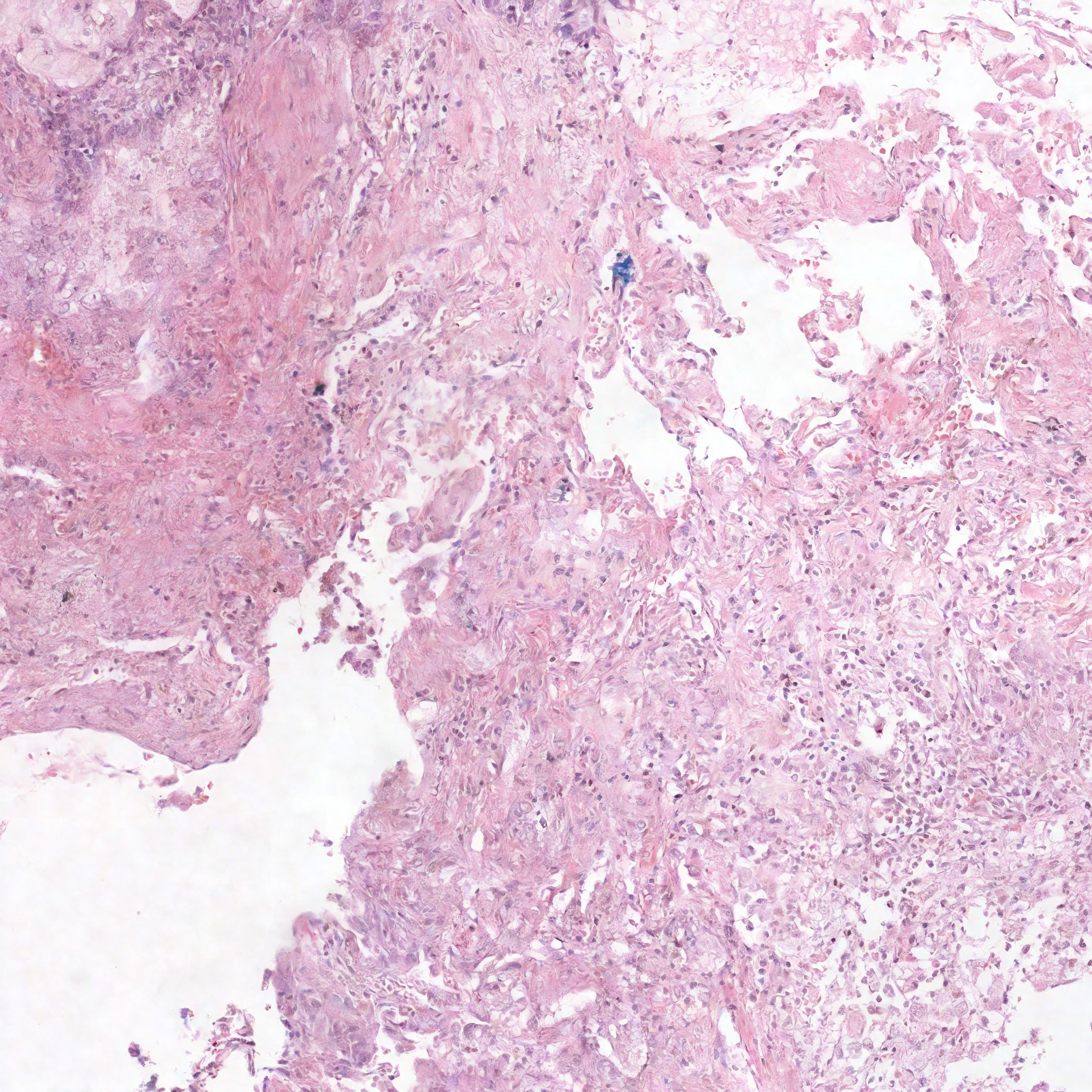
DiffInfinite conditioning control visualization. The images is is generated from the squared mask shown (2048x2048 pixels). This output doesn't have biological meaning because the generated class depends on the mask shape and its neighbours. The mask generation is necessary to give this spatial information.

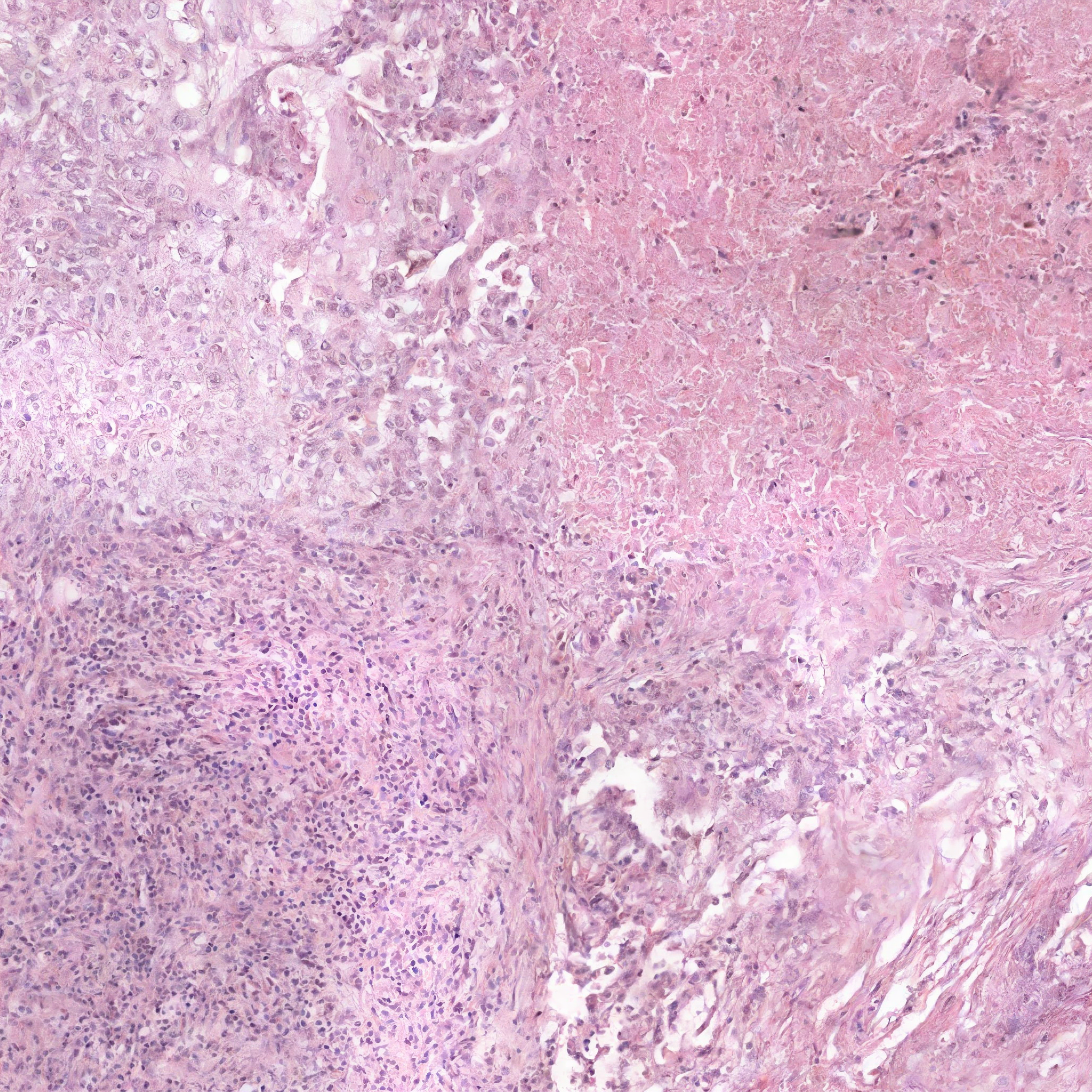
Generated images conditioned on the synthetic segmentation masks.
Unknown

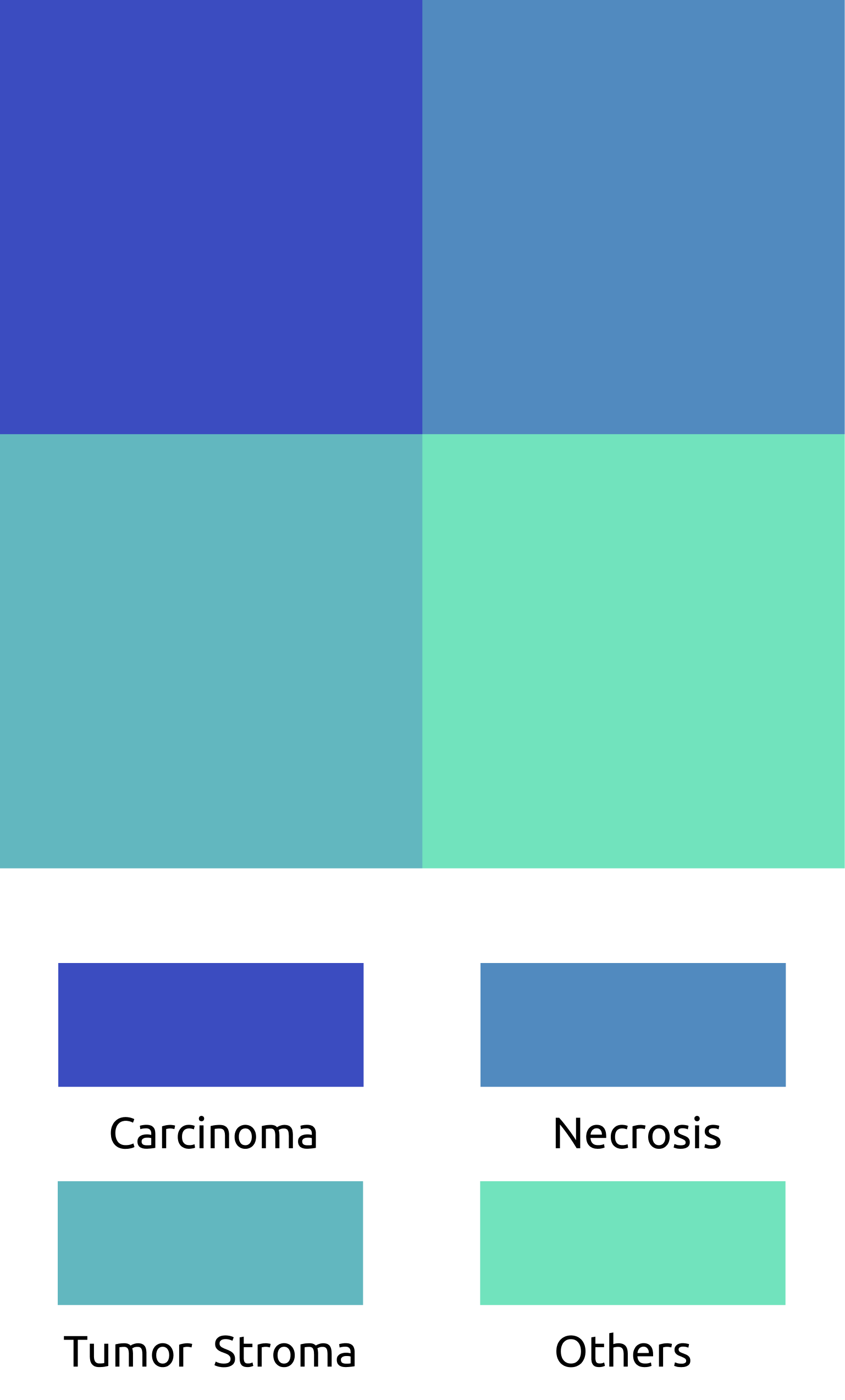
Carcinoma

Necrosis

Tumor Stroma

Others

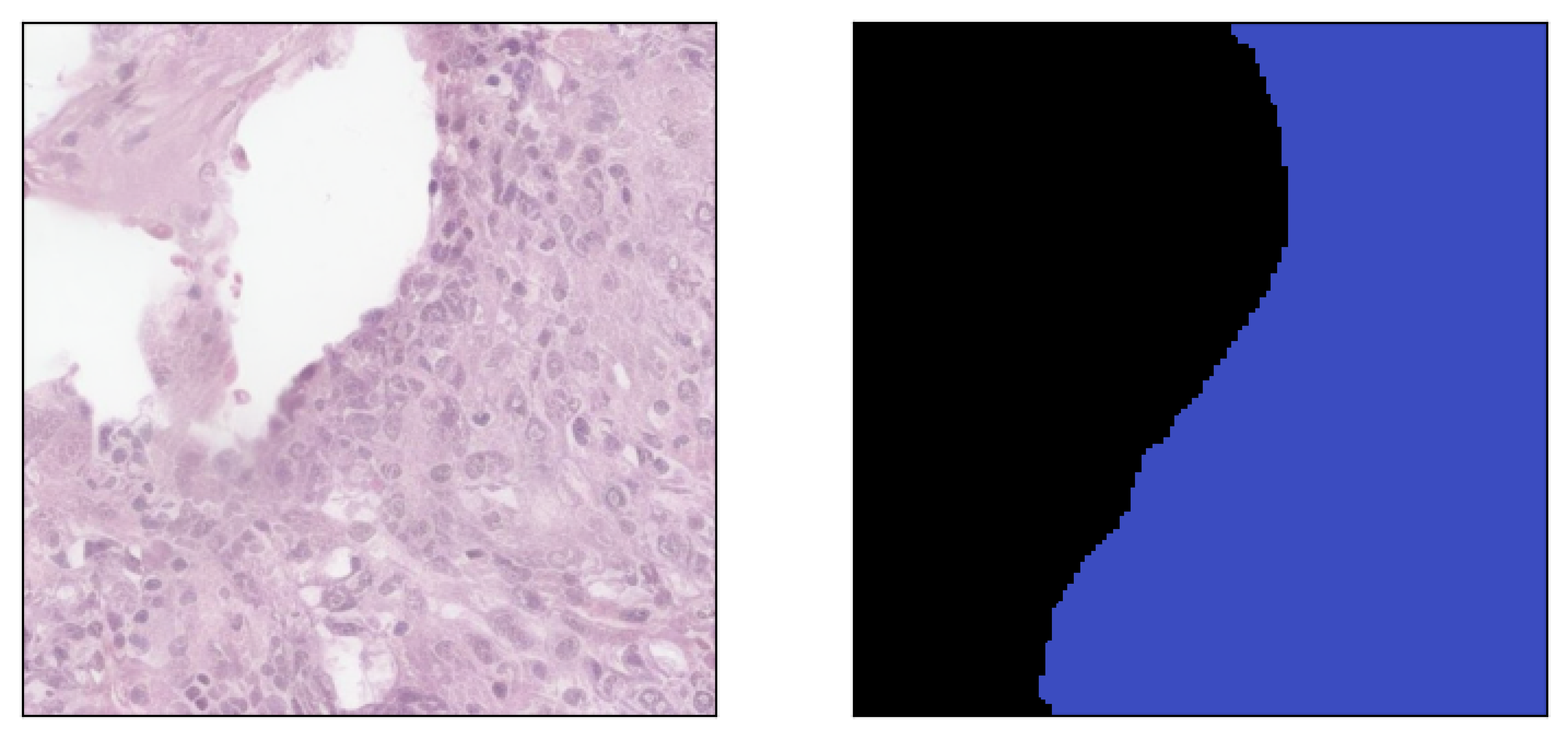
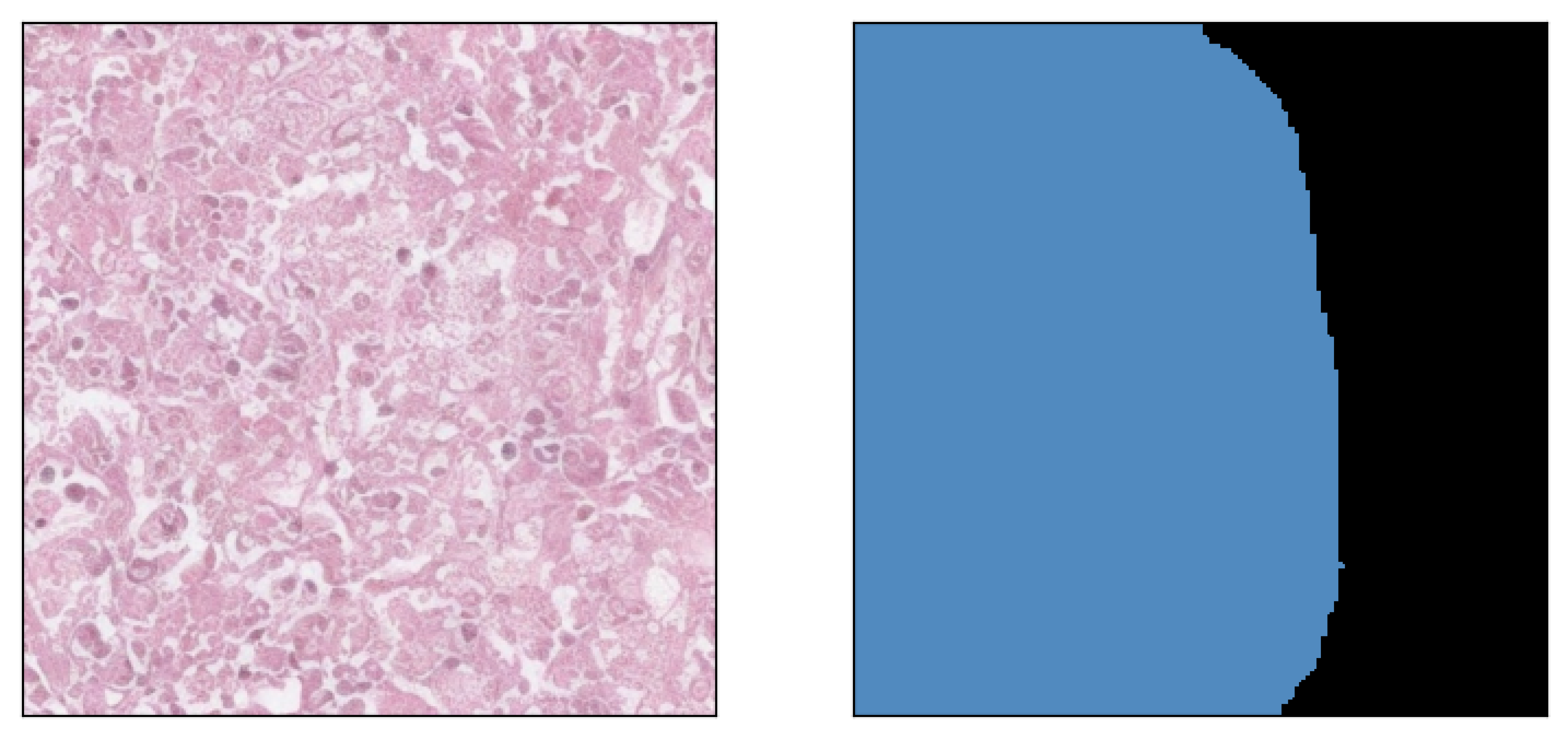
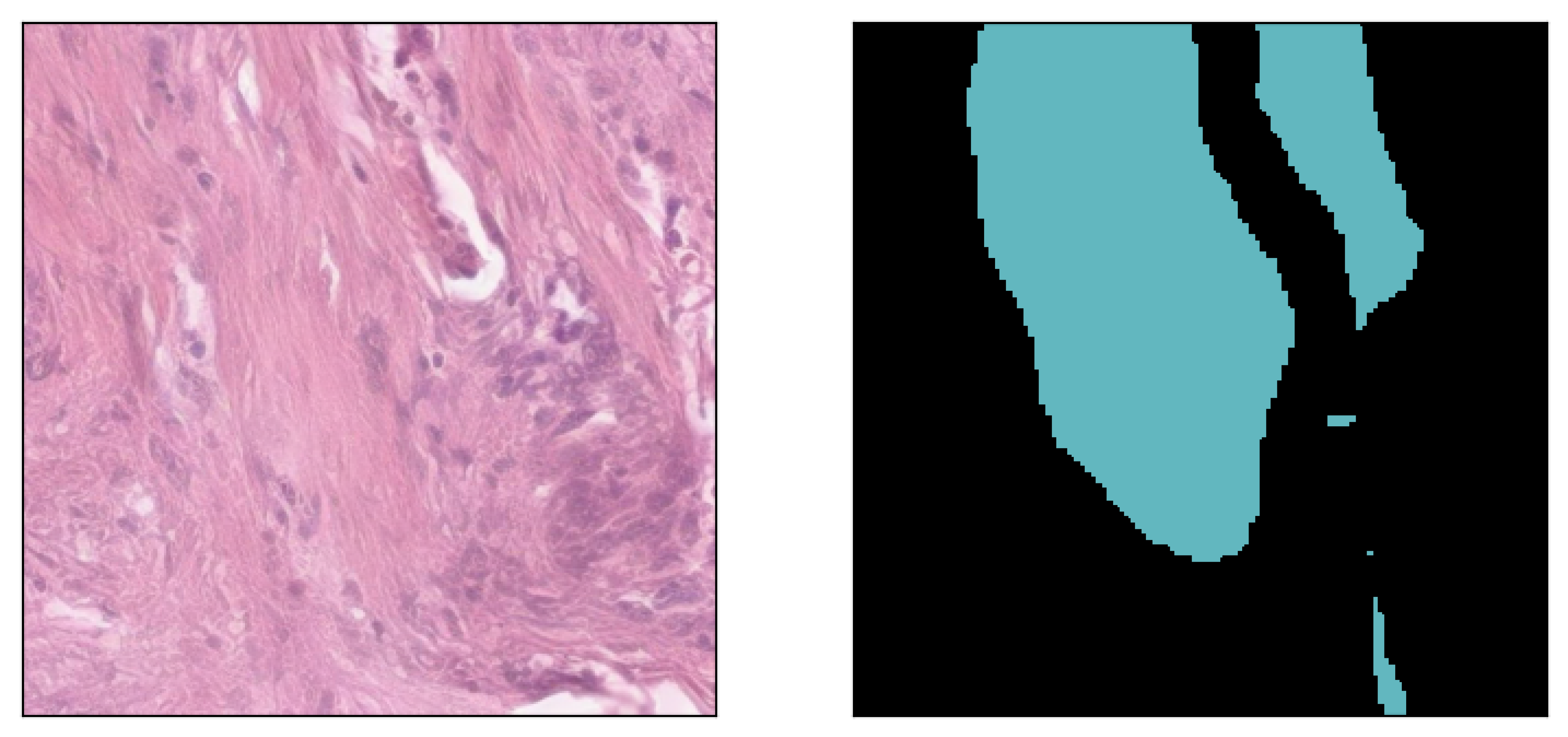
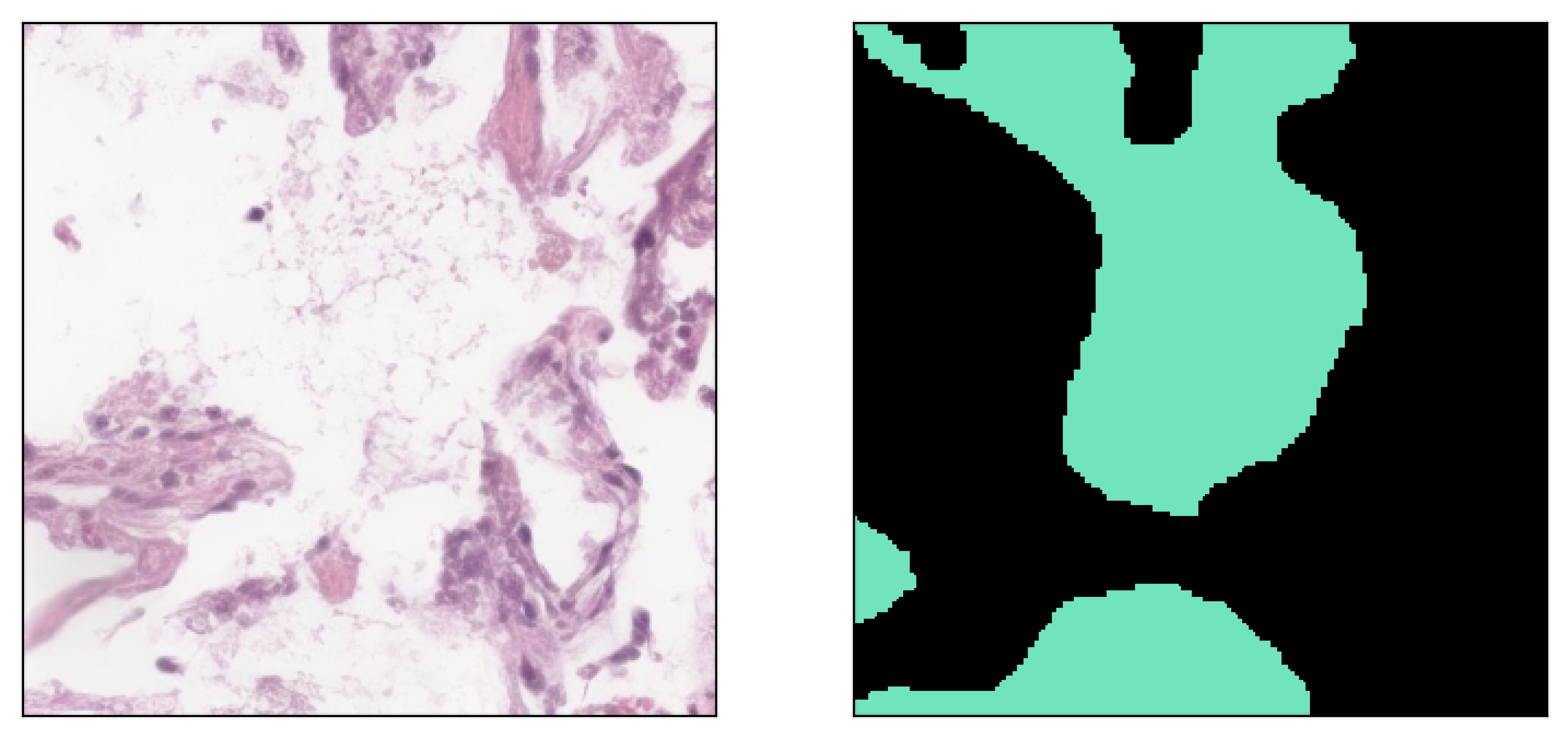
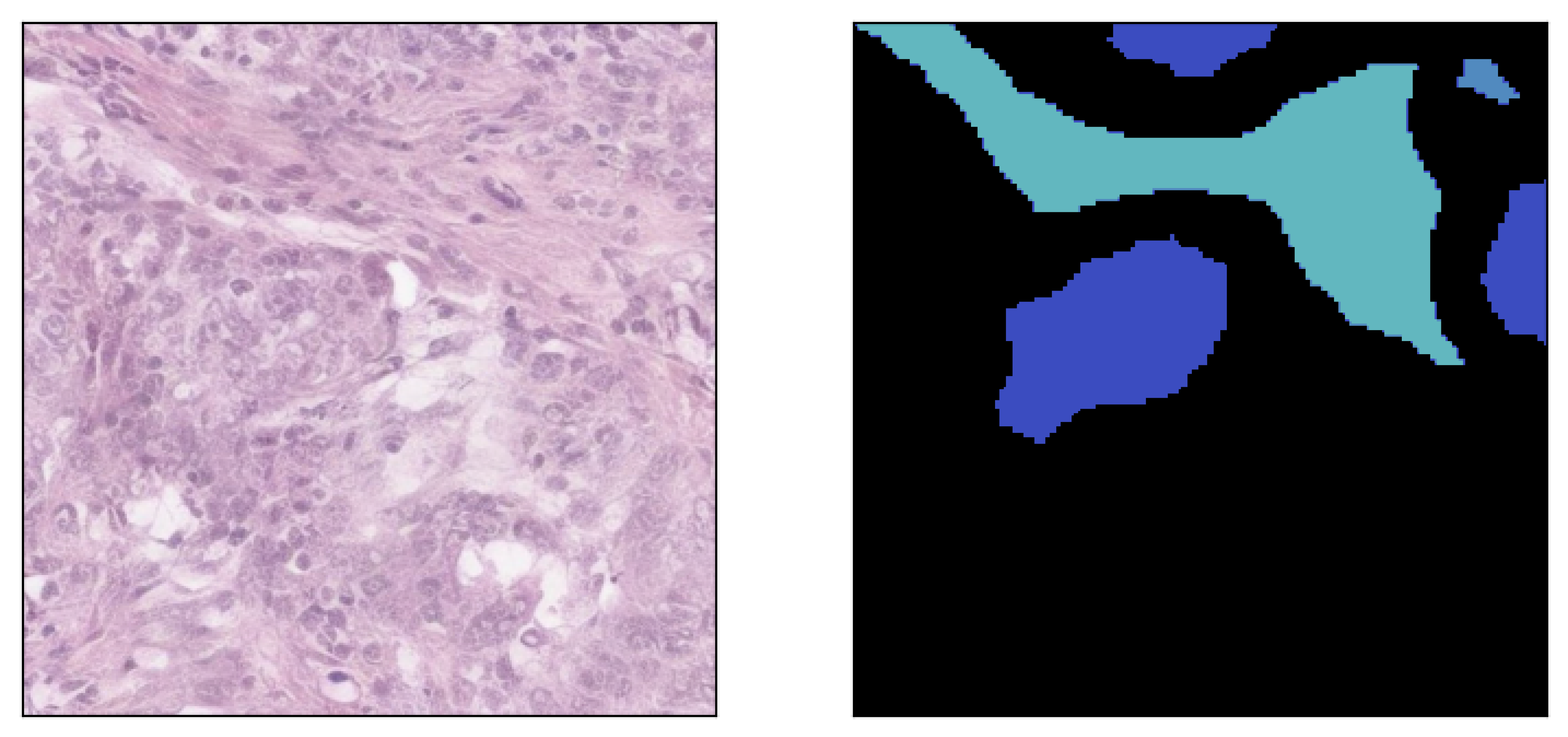
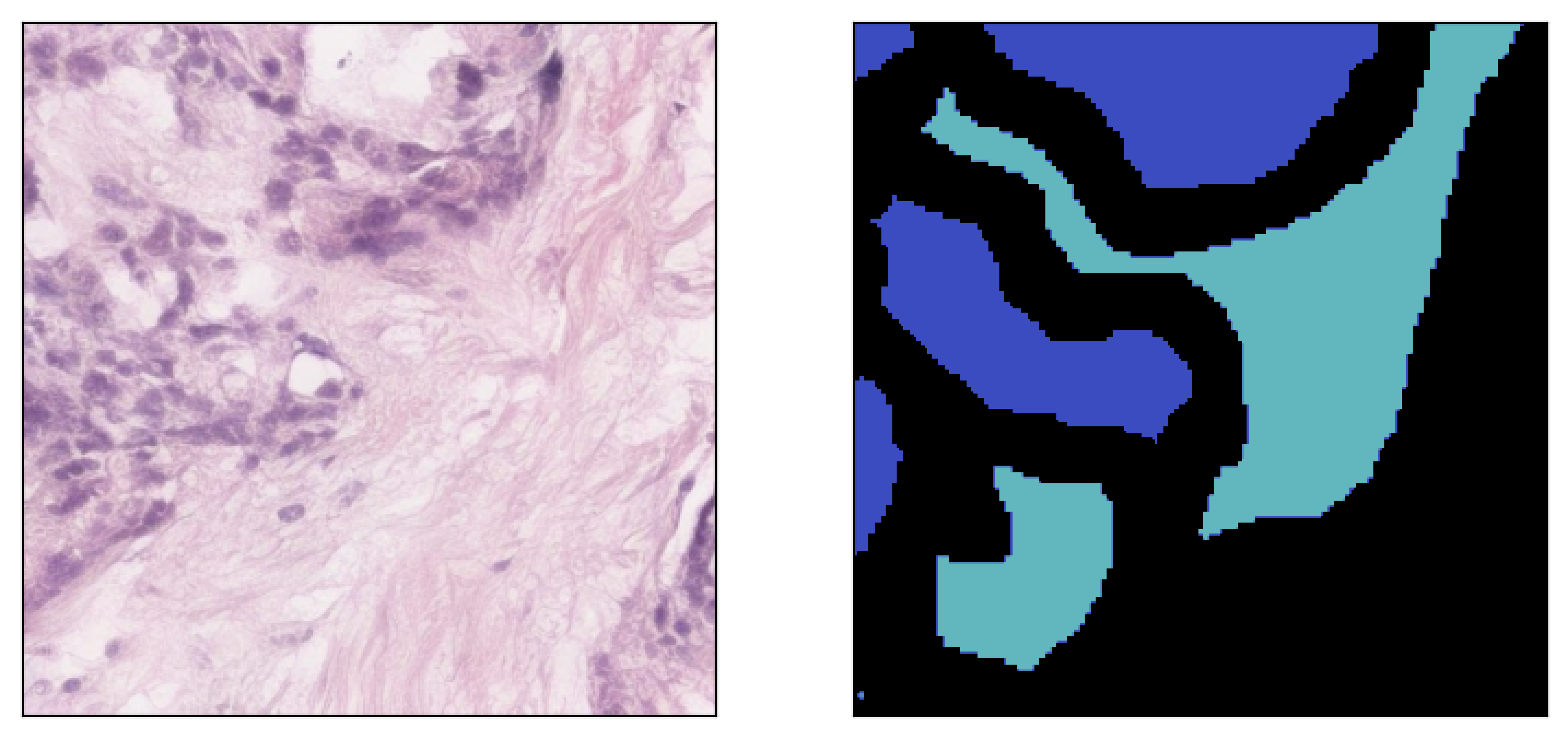
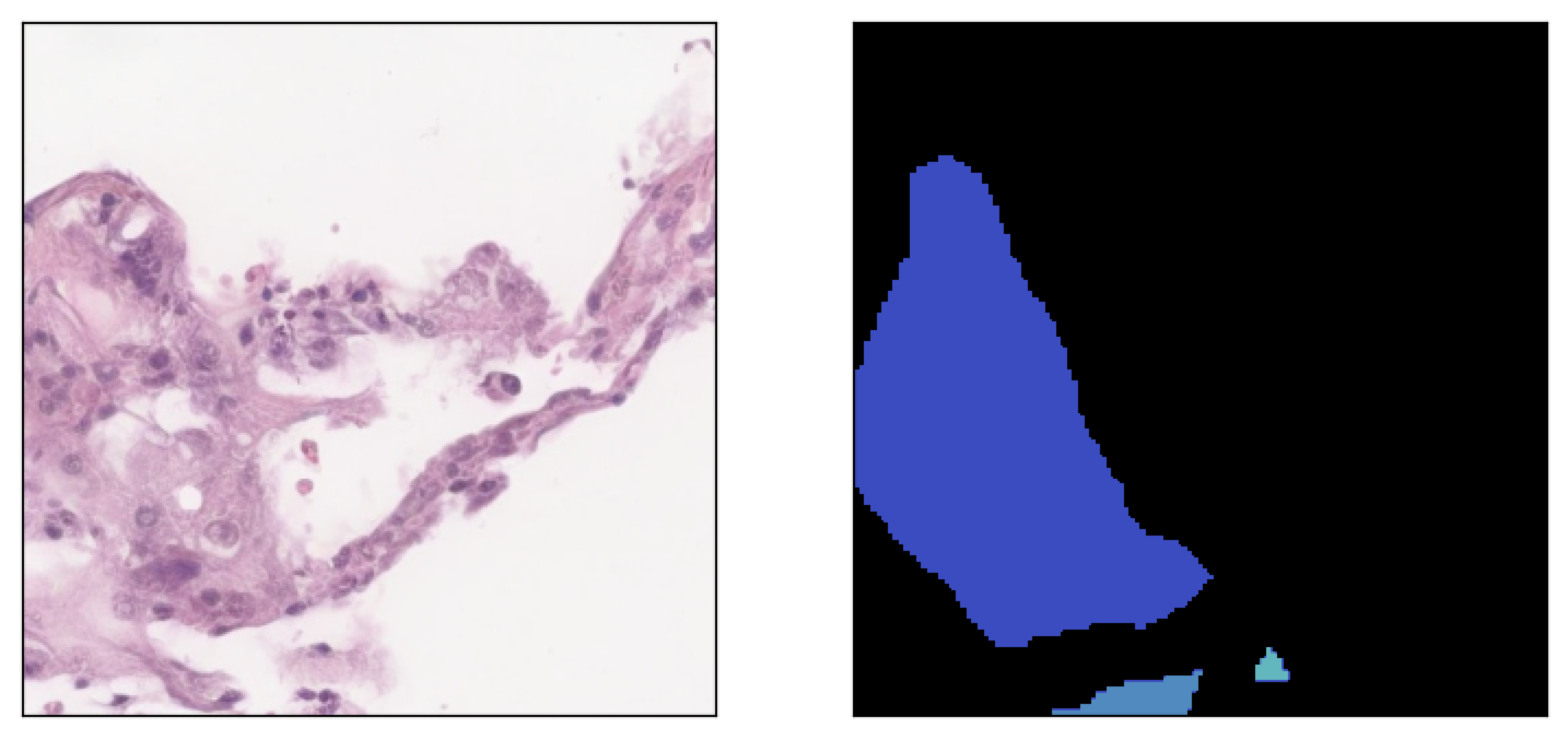
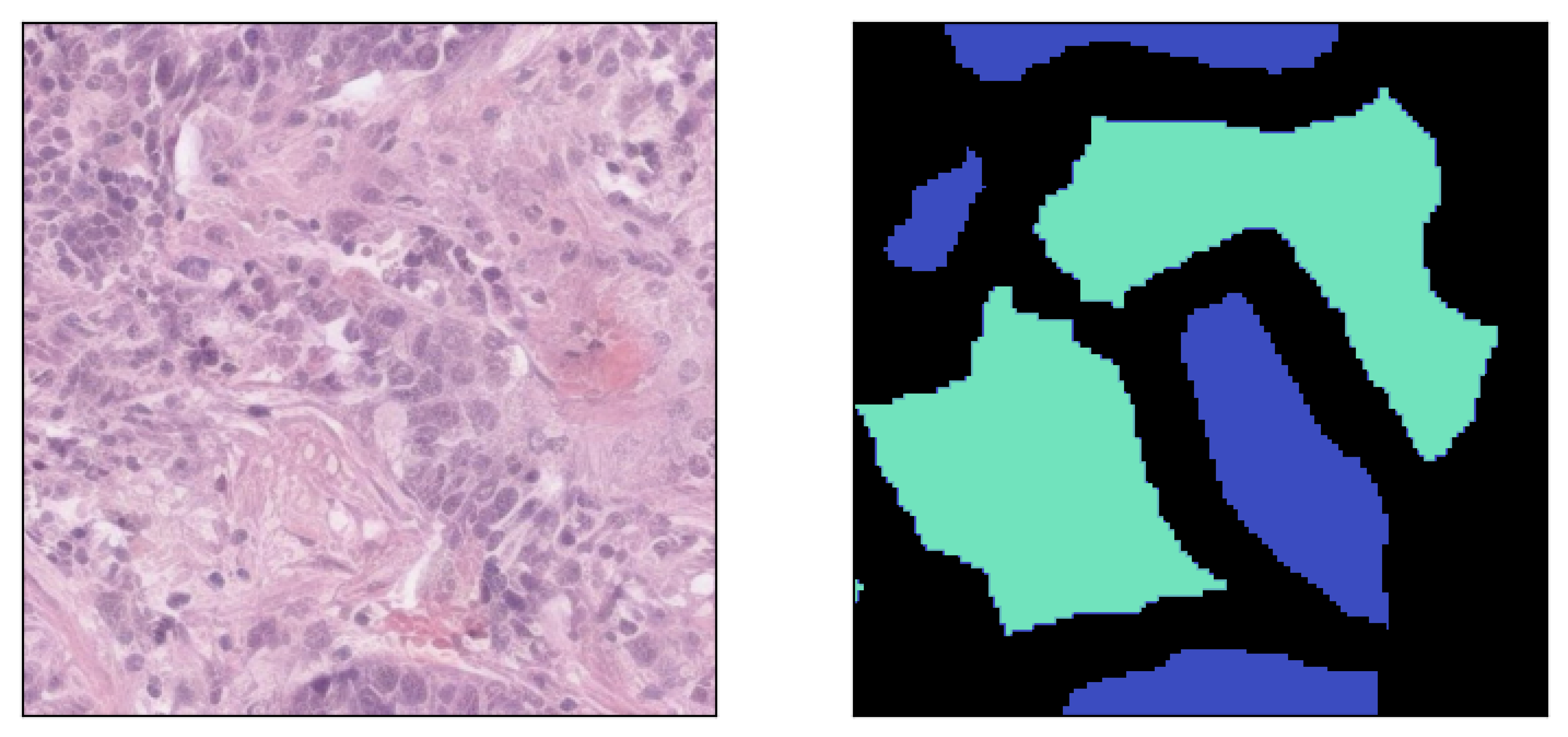
Image inpainted with different classes and different strengths of conditioning (small ω corresponding to less diversity)








@inproceedings{NEURIPS2023_f64927f5,
author = {Aversa, Marco and Nobis, Gabriel and H\"{a}gele, Miriam and Standvoss, Kai and Chirica, Mihaela and Murray-Smith, Roderick and Alaa, Ahmed M. and Ruff, Lukas and Ivanova, Daniela and Samek, Wojciech and Klauschen, Frederick and Sanguinetti, Bruno and Oala, Luis},
booktitle = {Advances in Neural Information Processing Systems},
editor = {A. Oh and T. Naumann and A. Globerson and K. Saenko and M. Hardt and S. Levine},
pages = {78126--78141},
publisher = {Curran Associates, Inc.},
title = {DiffInfinite: Large Mask-Image Synthesis via Parallel Random Patch Diffusion in Histopathology},
url = {https://proceedings.neurips.cc/paper_files/paper/2023/file/f64927f5de00c47899e6e58c731966b6-Paper-Datasets_and_Benchmarks.pdf},
volume = {36},
year = {2023}
}